How to construct a KO cell line
CRISPR/Cas9 technology enables the rapid generation of loss‐of‐function mutations in a targeted gene and changed the way of scientists approach their research. A single cell is a harbor of mutations that can be used to establish a new cell line, thereby creating a CRISPR‐induced knockout clone. These clonal cell lines serve as crucial tools for exploring protein function, analyzing the consequences of gene loss, and investigating the specificity of biological reagents. However, the successful derivation of knockout clones is technically challenging and in practice, it isn’t that simple. Obtaining high quality and reproducible editing results may need experience and optimization.
The standard workflow to generate CRISPR KO cell lines involves five major steps from project planning to confirmation of the desired edit. These steps are:
1. Design and production of KO guide
2. Cell transfection
3. Edited cells enrichments
4. Single-cell isolation and expansion
5. Confirmation of edits
Each of these steps requires careful consideration and technical skill in executing the many different methods involved.
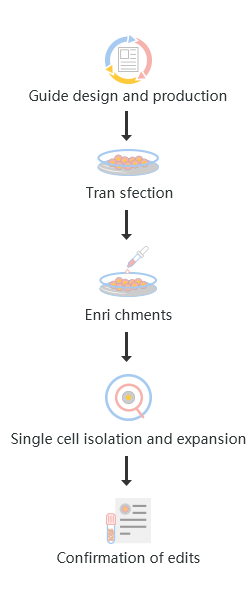
Fig : KO gene editing workflow
Guide design and production:
The guide RNA recognizes the target gene region and directs the gene-editing machinery to the target section of DNA. For gRNA production, it is required to design and synthesize gRNA oligo sequence. The synthesized oligo was then cloned into a suitable expression vector and verified the oligo sequence.
A researcher can use a variety of guide formats to generate KO cell lines; single guide RNA (sgRNA), a two-part oligo system of cr:tracrRNA, in vitro-transcribed RNA, plasmid, and lentivirus. The plasmids and lentivirus- based approach are most popular to researchers, though they are time-consuming, require selection agents, and prone to off-target edits. However, lentivirus may offer advantages for difficult-to-transfect cells. The use of sgRNAs offers reduced off-target effects and time-saving benefits. Furthermore, the use of multiple gRNAs can increase the chance of obtaining the desired edit and save time downstream in clone selection stages. But these gRNAs must be targeted correctly. To begin with, gRNA targeting the gene of interest, the gRNA needs to clone in a suitable expression vector and sequence verification of gRNA.
Red Cotton™ CRISPR Gene Editing Designer is a free tool for knockout cell line strategy design. It contains parameters of 800 cell lines, and combine with Ubigene’s advanced gRNA design principle. Simply enter your target gene, and you could get 3 different KO strategies. There is a Red Cotton™ gRNA plasmid bank that has the validated gRNAs in stock. It contains over 10,000 gRNA plasmids for knockout cell line generation. Each plasmid only takes 80 USD.
Guide design and production:
When the CRISPR KO strategy has been finalized, choose the most efficient method for cell transfection is next. Determining the most efficient method is considered the most difficult step in the CRISPR workflow by many researchers. Each cell type often required an extensive optimization of the transfection conditions to accurately deliver the CRISPR gene-editing machinery into the cell. Besides that, some cell types, including stem cells, primary cells and immune and hematopoietic cell lineages are more difficult to transfect, as they have altered cell repair mechanisms, and lower cell viability. Thus, a successful transfection often depends upon previous experience with the cell line of choice to understand the cell characteristics.
Ubigene has worked on over 100 tyes of cell lines, and had developed a mature procedure to explore the optimal cell transfection method.
Enrichments and single cell expansion:
A clonal selection workflow is considered the best practice to develop a homogenous cell population that contains the desired KO. The researcher usually uses two common methods for enrichments and single cell expansion, one is dilution cloning and another, fluorescence-activated cell sorting (FACS) of single cells. Although both approaches can yield single cell–derived clones. FACS sorting is often more effective given its ability to specifically isolate double-positive cells. However, dilution cloning is cheaper and may cause less cellular stress than sorting. Furthermore, in this stage researcher should consider fully optimized growth conditions, otherwise, it could lead to the death of precious cell samples and the loss of all work in generating KO cell lines.
For some cell lines, generating sigle-cell clone is tricky, only a few clones may be able to thrive during isolation. Ubigene has rich experience on clonal isolation, so preliminary tests would be done to find out the best way to generate single-cell clones.
Confirmations of edits:
Once a KO cell line has been established, adequate validation of the specific gene edits is necessary. This requires the characterization of the KO and ensure the downstream viability of the work. Multiple methods offer the best assurance of an accurate gene edit. Common methods to validate engineered cell lines include Sanger sequencing, next-generation sequencing, and qPCR to verify the edit at a genomic level. Western blot and mass spectrometry can confirm of the KO at the proteomic level. For functional studies immunohistochemistry, immunocytochemistry and FACS are often used.
Ubigene CRISPR-U™ KO cell line:
Ubigene developed the CRISPR-UTM system for precise genome editing and our KO cell lines have been extensively validated using qPCR, Sanger sequencing, and in many cases, Western blotting to confirm full gene ablation at the genomic and proteomic levels. By using CRISPR-U™ developed KO cell line, researcher can reduce their experimental time and achieve target solutions quicker. Now Ubigene offer the best KO cell line service ever! Only 3480 USD, over 5000 genes guarantee WB result negative.
Reference:
Generating Single Cell–Derived Knockout Clones in Mammalian Cells with CRISPR/Cas9. Curr Protoc Mol Biol. 2019.
 Subscribe Us
Subscribe Us Gene Editing Services
Gene Editing Services
 EZ-editor™
EZ-editor™ Red Cotton Gene knockout Project
Red Cotton Gene knockout Project









